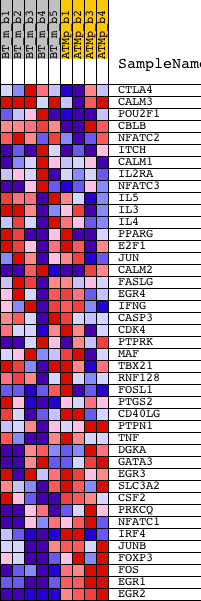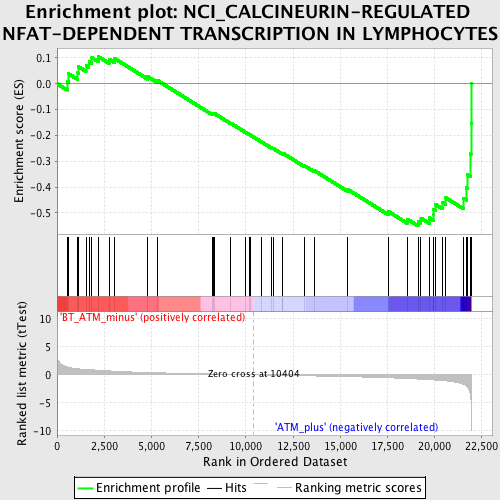
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | NCI_CALCINEURIN-REGULATED NFAT-DEPENDENT TRANSCRIPTION IN LYMPHOCYTES |
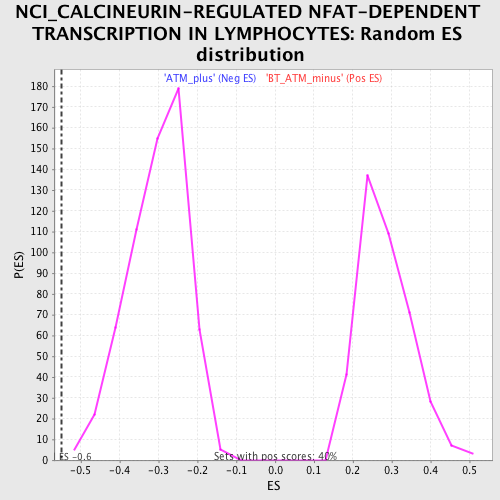
| Enrichment Score (ES) | -0.5503041 |
| Normalized Enrichment Score (NES) | -1.8182398 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.25414097 |
| FWER p-Value | 0.645 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CTLA4 | 1419334_at | 529 | 1.413 | 0.0100 | No | ||
| 2 | CALM3 | 1426710_at 1438825_at 1438826_x_at 1450864_at | 608 | 1.338 | 0.0388 | No | ||
| 3 | POU2F1 | 1419716_a_at 1427695_a_at 1427835_at | 1100 | 1.059 | 0.0420 | No | ||
| 4 | CBLB | 1437304_at 1455082_at 1458469_at | 1123 | 1.052 | 0.0664 | No | ||
| 5 | NFATC2 | 1425901_at 1425990_a_at 1426031_a_at 1426032_at 1439205_at 1440426_at | 1531 | 0.962 | 0.0711 | No | ||
| 6 | ITCH | 1415769_at 1431316_at 1459332_at | 1688 | 0.928 | 0.0864 | No | ||
| 7 | CALM1 | 1417365_a_at 1417366_s_at 1433592_at 1454611_a_at 1455571_x_at | 1843 | 0.892 | 0.1009 | No | ||
| 8 | IL2RA | 1420691_at 1420692_at | 2200 | 0.808 | 0.1042 | No | ||
| 9 | NFATC3 | 1419976_s_at 1452497_a_at | 2750 | 0.690 | 0.0958 | No | ||
| 10 | IL5 | 1425546_a_at 1442049_at 1450550_at | 3035 | 0.639 | 0.0983 | No | ||
| 11 | IL3 | 1450566_at | 4802 | 0.410 | 0.0275 | No | ||
| 12 | IL4 | 1449864_at | 5329 | 0.360 | 0.0122 | No | ||
| 13 | PPARG | 1420715_a_at | 8208 | 0.139 | -0.1159 | No | ||
| 14 | E2F1 | 1417878_at 1431875_a_at 1447840_x_at | 8294 | 0.134 | -0.1165 | No | ||
| 15 | JUN | 1417409_at 1448694_at | 8322 | 0.132 | -0.1146 | No | ||
| 16 | CALM2 | 1422414_a_at 1423807_a_at | 9204 | 0.077 | -0.1530 | No | ||
| 17 | FASLG | 1418803_a_at 1449235_at | 9986 | 0.027 | -0.1880 | No | ||
| 18 | EGR4 | 1449977_at | 10167 | 0.015 | -0.1959 | No | ||
| 19 | IFNG | 1425947_at | 10234 | 0.011 | -0.1986 | No | ||
| 20 | CASP3 | 1426165_a_at 1430192_at 1449839_at | 10836 | -0.027 | -0.2254 | No | ||
| 21 | CDK4 | 1422441_x_at | 11336 | -0.058 | -0.2468 | No | ||
| 22 | PTPRK | 1423277_at 1423278_at 1431680_a_at 1441276_at 1441277_s_at 1441644_at 1443491_at 1443699_at 1444763_at | 11448 | -0.065 | -0.2503 | No | ||
| 23 | MAF | 1435828_at 1437473_at 1444073_at 1447849_s_at 1447945_at 1456060_at | 11924 | -0.094 | -0.2697 | No | ||
| 24 | TBX21 | 1449361_at | 11939 | -0.095 | -0.2680 | No | ||
| 25 | RNF128 | 1418318_at 1449036_at | 13088 | -0.166 | -0.3165 | No | ||
| 26 | FOSL1 | 1417487_at 1417488_at | 13631 | -0.203 | -0.3363 | No | ||
| 27 | PTGS2 | 1417262_at 1417263_at | 15377 | -0.327 | -0.4081 | No | ||
| 28 | CD40LG | 1422283_at | 17543 | -0.526 | -0.4943 | No | ||
| 29 | PTPN1 | 1417068_a_at 1438670_at | 18535 | -0.658 | -0.5236 | No | ||
| 30 | TNF | 1419607_at | 19120 | -0.750 | -0.5322 | Yes | ||
| 31 | DGKA | 1418578_at | 19270 | -0.779 | -0.5201 | Yes | ||
| 32 | GATA3 | 1448886_at | 19704 | -0.862 | -0.5191 | Yes | ||
| 33 | EGR3 | 1421486_at 1436329_at | 19914 | -0.912 | -0.5066 | Yes | ||
| 34 | SLC3A2 | 1425364_a_at | 19929 | -0.915 | -0.4851 | Yes | ||
| 35 | CSF2 | 1427429_at | 20032 | -0.940 | -0.4670 | Yes | ||
| 36 | PRKCQ | 1426044_a_at | 20416 | -1.029 | -0.4596 | Yes | ||
| 37 | NFATC1 | 1417621_at 1425761_a_at 1428479_at 1447084_at 1447085_s_at | 20589 | -1.073 | -0.4415 | Yes | ||
| 38 | IRF4 | 1421173_at 1421174_at | 21520 | -1.685 | -0.4433 | Yes | ||
| 39 | JUNB | 1415899_at | 21672 | -1.992 | -0.4020 | Yes | ||
| 40 | FOXP3 | 1420765_a_at 1457082_at | 21728 | -2.176 | -0.3519 | Yes | ||
| 41 | FOS | 1423100_at | 21901 | -3.644 | -0.2716 | Yes | ||
| 42 | EGR1 | 1417065_at | 21922 | -4.993 | -0.1518 | Yes | ||
| 43 | EGR2 | 1427682_a_at 1427683_at | 21930 | -6.298 | 0.0002 | Yes |